Wildlife Genomics
Wildlife Genomics
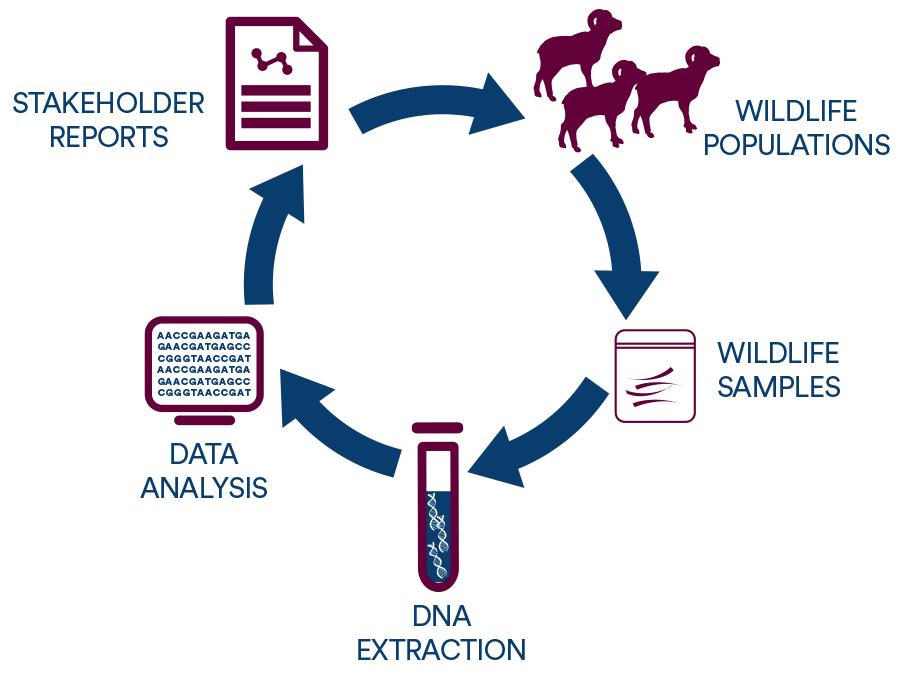
We use tissue, blood, horn, hair, and even fecal samples from wildlife populations to extract DNA. We then analyze the genetic information from DNA to learn about population history, health, and dynamics. We share that information with stakeholders in wildlife conservation, from citizens to state and federal wildlife agencies, so that they can make informed decisions about wildlife management.
Wildlife genomics is the application of large-scale DNA tools on wildlife for conservation. Our lab works on a variety of wildlife species with the goal of contributing to effective conservation and management. Genetic information can be used by wildlife managers and conservationists to determine conservation units, calculate sustainable harvest rates, and manage translocation.
Most wildlife genetic work to date has involved using a handful of DNA markers, from 10 to 100. A DNA marker is a place in the genome. For example, single DNA base (the chemicals called A,C,G, and T), a repeated sequence of bases, or a sequence of a gene are examples of different kinds of DNA markers. Genomic tools scale up the number of markers to thousands of genetic markers all the way up to whole genome sequences.
Using high numbers of markers enables us to learn vital information for wildlife conservation. First, using many genetic markers allows us to address traditional questions in conservation and population genetics with greater accuracy. This includes questions about population structure, genetic diversity, gene flow, and inbreeding. Gene flow happens when new or migrating individuals mate with local populations, thereby transferring genetic material between populations.
Second, using genomics allows us to address new questions about the actual genes involved in local adaptation, health, behavior, and inbreeding depression. Local adaptation is how well an organism matches its specific environmental. Inbreeding depression is the reduction in the ability to survive and reproduce as a result of mating between close relatives. Knowing the specific genes involved in traits such as disease resistance or migration can help predict the effect of climate change and land use decisions on wildlife.
Learn more about wildlife genomics:

